A Novel Machine Learning Based Screening Method For High-Risk Covid-19 Patients Based On Simple Blood Exams
This paper presents a predictive model to potentially identify high-risk COVID-19 infected patients based on easily analyzed circulatory blood markers. These findings can enable effective and efficient care programs for high-risk patients and periodic monitoring for the low-risk ones, thereby easing the hospital flow of patients and can further be utilized for hospital bed utilization assessment. The present machine learning-based SV-LAR model results in a high 87% f1 score, harmonic mean of 91% precision, and 83% recall to classify COVID-19, infected patients, as high-risk patients needing hospitalization.
COVID-19 is a new disease for which effective treatment is still awaited. It was declared a pandemic by World Health Organization (WHO) on March 11, 2020. As of January 28, 2021, more than 100 million people have been affected by this infection causing more than 2 million fatalities. Global health care now faces unprecedented challenges with the widespread and rapid human-to-human transmission of SARS-CoV-2 and high morbidity and mortality with COVID-19 worldwide. COVID-19 patients get worse quickly and aggressively. In addition to high transmissibility SARS-CoV-2 infection it is also characterized by fever, dry cough, weakness, headache, dyspnoea, and loss of smell and taste in the early stages, which are common symptom of cold and flu. The early onset of common symptoms can rapidly change to acute respiratory distress syndrome (ARDS), acute cardiac injury, cytokine storm, coagulation dysfunction, and multi-organ failure if the disease is not resolved, resulting in patient death.
Early studies showed that COVID-19 patients with comorbidity may lead to poor prognosis, increasing the risk of severe illness from COVID-19. Among laboratory-confirmed cases, patients with any comorbidity yielded poorer clinical outcomes than those without. Several studies have been conducted to find a correlation between pre-existing medical conditions and their impact on COVID-19 prognosis. In a meta-analysis by Wang et al, Hypertension, diabetes, Chronic obstructive pulmonary disease (COPD), cardiovascular disease, and cerebrovascular disease were found to be the major risk factors for patients with COVID-19. Several risk factors that led to the progression of COVID-19 pneumonia were identified, including age, history of smoking, maximum body temperature at admission, respiratory failure, albumin, and C-reactive protein. Given the virtually unstoppable global trend of SARS-CoV-2, together with the high prevalence of comorbidities worldwide, the combination of these two conditions poses greater clinical, societal, and economic burdens to healthcare systems
Until now the source of the pathogenesis of the COVID-19 remains unclear, and no specific treatment has been recommended for coronavirus infection except for meticulous care. The world is ready to receive the vaccines as approved worldwide, but the threat continues with mutating strains of the virus. Therefore, the need for a better solution for providing care to those who absolutely need it and to predict the future requirements for better planning and management for better patient outcomes, continues. In several articles, researches have indicated the need for better hospital management by early identification of patients requiring hospitalization and possible further triage.
In another attempt to decode the comorbidity-related risks in COVID-19 patients, Zhao et al. developed a logistic regression-based classification model to predict two primary outcomes of admission to the intensive care unit (ICU) and death. The risk score model yielded accuracy with an Area under the curve (AUC) of 0.74 ([95% CI, 0.63-0.85], $p=0.001$) for predicting ICU admission and 0.83 ([95% CI, 0.73-0.92], $p<0.001$) for predicting mortality for the testing dataset. This model was developed and internally validated using data from the COVID-19 persons under investigation (PUI) registry of 4997 patients from a major academic hospital (Stony Brook University Hospital) in New York. Another finding was that the mortality group uniquely contained cardiopulmonary parameters as top predictors.
In another study aimed to clarify high-risk factors for COVID-19, researchers used Multivariate Cox regression to identify risk factors associated with the progression of the disease. Univariate and multivariate analyses showed that comorbidity, older age, lower lymphocyte count, and higher lactate dehydrogenase at presentation were independent high-risk factors for progression. A novel scoring model, named CALL, with an area under the receiver operating characteristic curve (ROC) of 0.91 (95% CI, .86-.94) was established to help clinicians better choose a therapeutic strategy. Elisa Grifoni et al, tested the predictive power of the CALL score in an Italian COVID-19 population admitted to hospital from 12 March to 20 April 2020 and consisting of 210 patients. Their findings concluded that the CALL score is a good prognosticator for in-hospital mortality but not for progression to severe COVID-19 in their settings.
In another statistical analysis regarding the associations between increased cardiac injury markers and the risk of 28-day-all-cause death of COVID-19 patients in the Chinese population, the 5 myocardial biomarkers (high-sensitivity cardiac troponin I, creatine phosphokinase)-MB, N-terminal pro-B-type natriuretic peptide, creatine phosphokinase, and myoglobin) were found to be significantly prognostic of COVID-19 mortality. Baseline patient characteristics, laboratory markers, and chest radiography can predict short-term critical illness in hospitalized patients with COVID-19, with an internally validated $AUC=0.77$ using a logistic regression-based risk model developed by Steven Schalekamp, et al. In another study, Zhou et al. validated a nomogram including 6 predictors: age, respiratory rate, systolic blood pressure, smoking status, fever, and chronic kidney disease. The model demonstrated a high discriminative ability in the training cohort $(C-index=0.829)$, which was confirmed in the external validation cohort $(C-index=0.776)$ In addition, the calibration plots confirmed good concordance for predicting the risk of ICU admission.
In another study, a regression analysis showed that C-reactive protein (CRP) was significantly associated with aggravation of non-severe COVID-19 patients, with an area under the curve of 0.844 (95% confidence interval, 0.761-0.926) and an optimal threshold value of $26.9~mg/L$. In a Spanish study, COVID-19 patients with normal levels of lymphocytes or mild lymphopenia, imbalanced lymphocyte subpopulations were early markers of in-hospital mortality.
Despite several initiatives aimed at containing the spread of the disease, countries are faced with unmanageable increases in the demand for hospitalization and ICU beds. The health care system globally, has been stressed and stretched to its limit. In order to help in patient triage, several attempts have been made to discover early predictors of COVID-19 disease progression and spread. Identification of such factors that predict complications of COVID-19 is pivotal for guiding clinical care, improving patient outcomes, and allocating scarce resources effectively in a pandemic. Medical resource allocation assessments should be based on a risk/benefit approach considering the intensity of transmission, the health system's capacity to respond, other contextual considerations (such as upcoming events which may alter transmission or capacity) and the overall strategic approach to responding to COVID-19 in each specific setting. We think that it requires agile decision-making based on ongoing situational assessments at the most local administrative level possible.
We propose a predictive machine learning model that identifies a potential high-risk patient from the COVID-19 patient population based on blood based circulatory markers. These predictions would help the administrators to make provisions for the scarce hospital beds. Consequently, the model can help in providing better public health and social measures to alleviate patient care during the pandemic time thereby improving patient outcomes at large.
A. The Dataset
We have used data published on a public forum, that of Hospital Israelita Albert Einstein, at São Paulo, Brazil. The dataset contains records of patients that were tested for COVID-19 using SARS-CoV-2 Reverse transcription polymerase chain reaction (RT-PCR) and additional blood tests between the 28th of March 2020 and 3rd of April 2020. All data were anonymized following the best international practices and recommendations. The full dataset released included 5,644 individual patients' clinical test results that were standardized to have a mean of zero and a unit standard deviation. It provided information of patient hospitalization into three types of wards in the hospital, such as regular ward, semi-intensive care unit, and intensive care unit as depicted in Fig 1.
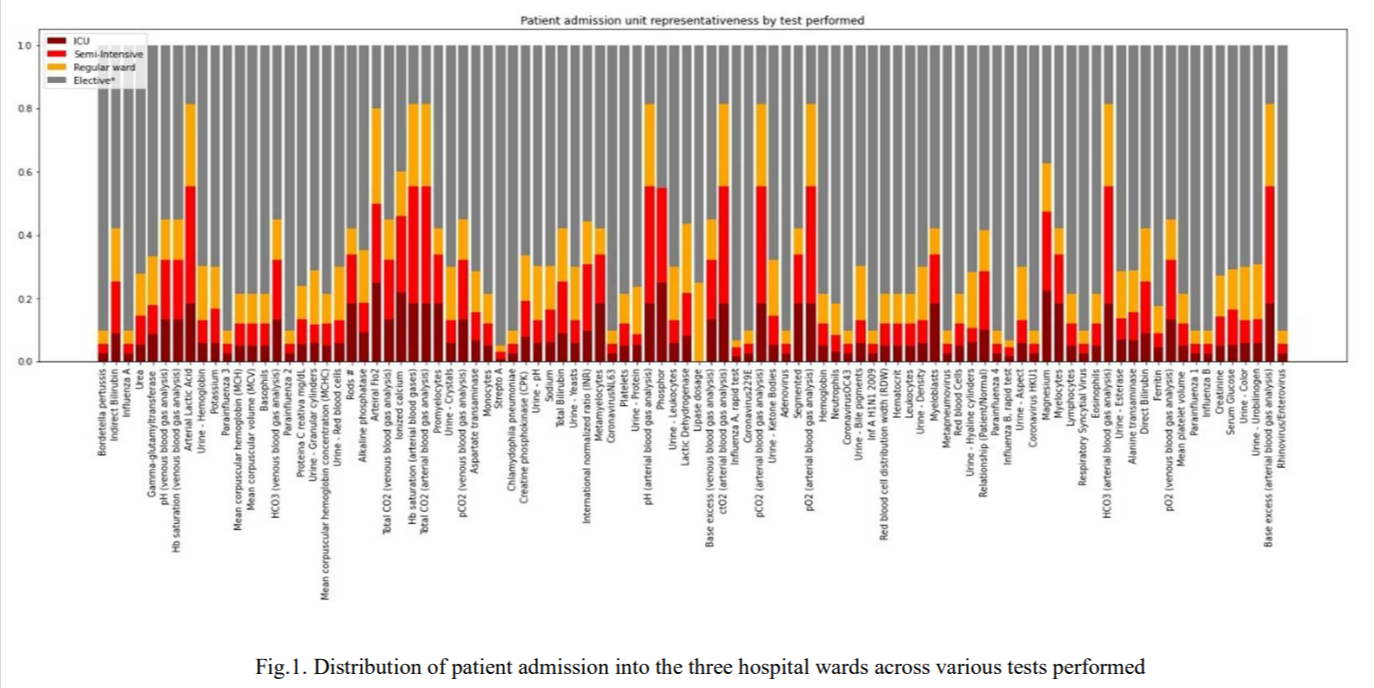
The information of patients admission to various wards in the hospital was used to create the target variable for the current problem statement. Hospitalization is needed by patients needing extra care and monitoring due to health who despite being infected do not need hospital admission constitute the low-risk patient population. This formed the basis of the binary classification and the target label for classification of patients needing hospitalization in any of the hospital wards $=1$, no hospitalization needed $=0$ for the current objective. The distribution as per figure 1 above was therefore transformed to look like Fig 2 below.

As the current hypothesis is set around blood analysis, we have carefully selected features of routine blood analysis only. Parameter related to patient age was not considered in order to avoid any age related bias in the present analysis. Tests pertaining to viral or bacterial infections other than SARS-CoV-2 were also dropped. It is our objective to find blood based markers to identify high-risk patients and therefore features related to routine urine analysis were also dropped from the current model. Blood gas analysis either on venous blood or arterial blood is also not included in the current analysis. Largely because the blood samples are required to be tested in a 30 minute window or need a cold supply chain. It is our intent to find markers that eases the hospital workload during the pandemic and therefore it is counterintuitive to include tests that need immediate attention and hospital setting to give good results. It is for this reason that the working dataset for the model building exercise includes test parameters form a simple blood workup, keeping in mind that the sample could be collected form patients home environment and not necessarily in the hospital setting. Fig 3 presents the frequency of each test performed amongst the blood analysis related parameters considered for the model building, in the select dataset of 558 patient records tested positive for SARS-CoV-2.

For the purposes of our research we extracted records of patients that were tested positive for the RT-PCR test for the ongoing COVID-19 infection. Our working dataset consisted 558 records of patients infected with COVID-19 from the whole of 5644 patient records. The target variable is the hospital admission class amongst the extracted records. Following our research question we formulated the hypothesis that we were to test -
H0: There is no correlation between blood analysis of a COVID positive patient and his/her hospitalization
H1: There is a correlation between blood analysis of a COVID positive patient and his/her hospitalization
B. Model building
We started model building after data processing. The working data had missing values and columns with all null values. Features with more than 95% missing values were dropped and remaining missing values were imputed with the mean. We started with simple linear algorithms such as logistic regression, ridge classification and elastic net, and moved on to non-parametric algorithm such as Nearest Neighbours Classifier and Gaussian Naive Bayes classification. We also used tree based algorithms like decision tree classifier and extra tree classifier. Multiple ensemble techniques like random forest classifier, bagging classifier, adaboost classifier were also used to model the target label with select features of the prepared dataset. We used scikit learn library of machine learning algorithms.
Based on our findings, we propose the SV-LAR model for our 2-class (SARS-CoV-2 positive induced hospitalization or not) classification. The proposed model uses voting classifier ensemble based on logistic regression, random forest and adaboost classifier. The working dataset also has class imbalance. Only 10% labels of the working dataset are positive class of hospitalised COVID positive patients, in the three available hospital wards. We have used SMOTE (Synthetic Minority Oversampling Technique) on the training data to deal with the class imbalance by upsampling the positive class.
C. Model performance measures
The performance of the model is expressed in terms Fl score, precision and recall. As we attempt imbalanced classification problem F1 score metric becomes more relevant. It is a measurement that considers both precision and recall to compute the score that can be interpreted as a weighted average of the precision and recall values. High F1 score (closer to 1.0) is desirable in our model. Precision is determined by the number of correctly labeled annotations divided by the total number of annotations added by the machine-learning annotator. It indicates how accurately the model has labelled the two classes. Another metric, recall specifies how many mentions that should have been annotated by a given label were actually annotated with that label. A recall score of 1.0 means that every mention that should have been labeled as entity type A was labeled correctly. In this imbalanced healthcare dataset high scores for precision and recall are desirable for an ultimate high fl score.
A. Evaluation
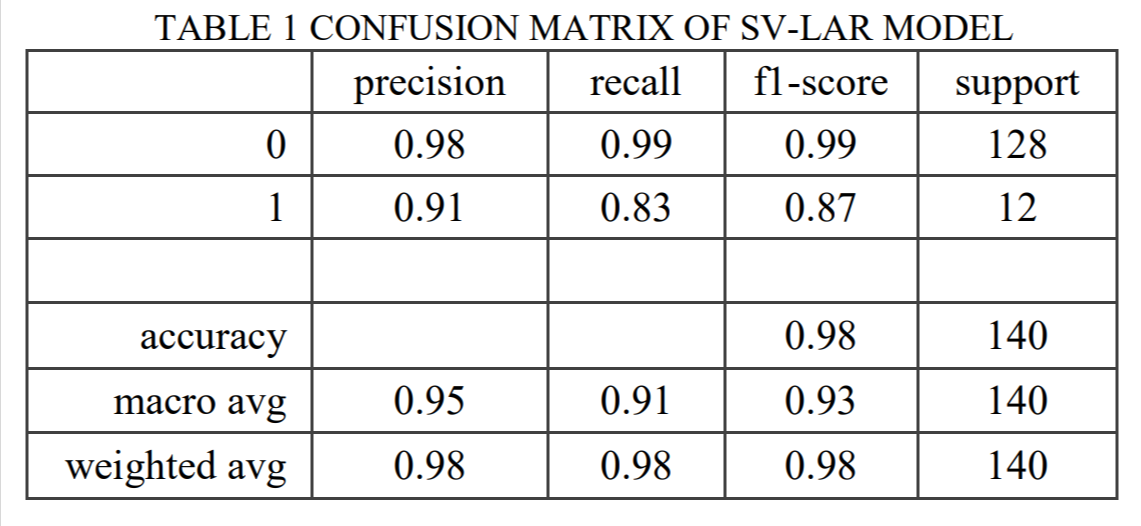
The proposed SV-LAR model of the COVID-19 infected patients produces a classification f1-score of 87%. With precision at 91% and recall at 83%. The confusion matrix is presented in the figure below.
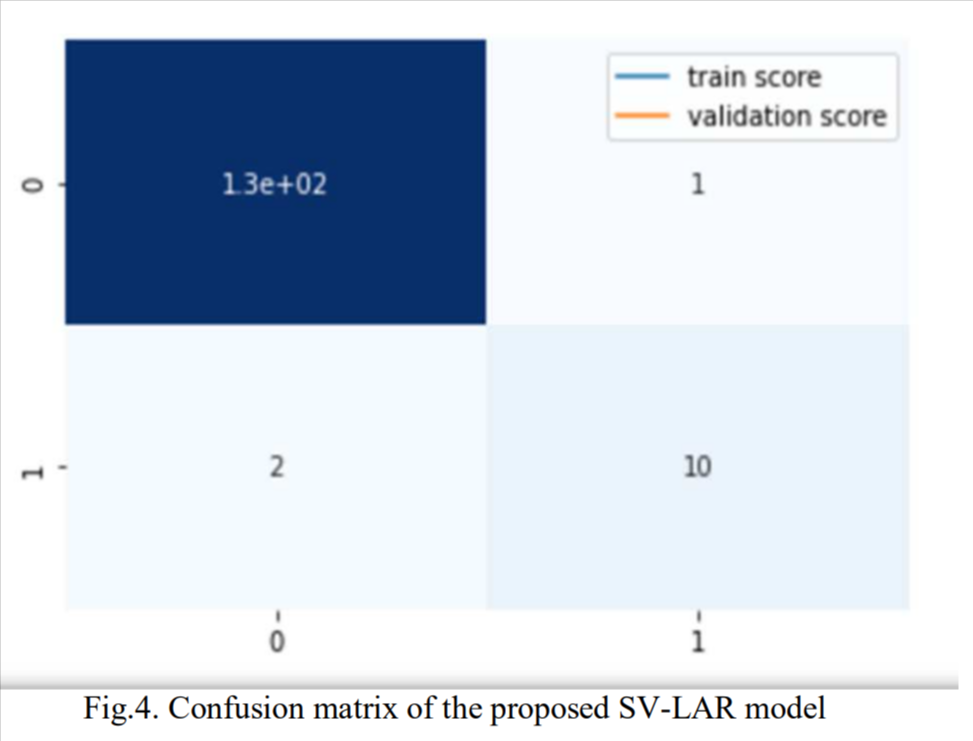
To the best of the authors knowledge, it is the first study to report a predictive machine learning model with high precision and recall for the triage of high-risk COVID-19 patients using simple blood exams. As per our findings, it would be possible to identify high-risk COVID-19 patients with more than 83% sensitivity and 91% specificity based on blood analysis alone. While the majority patients are asymptomatic, about 10% patients need hospitalisation. Our solution can enable healthcare professional, segregate potential high-risk patient in need of high degree of hospital care based on simple and cheap blood analysis and monitor them closely.
B. Proposed solution for patient management
Once our model is deployed in healthcare setting it will enable quick movement of patients and help manage hospital resources more effectively. The suspected patients take a SARS-CoV2-RT-PCR test and simultaneously give blood samples for analysis. If tested positive for COVID-19 infection their blood result is tested through the pretrained SV-LAR model.
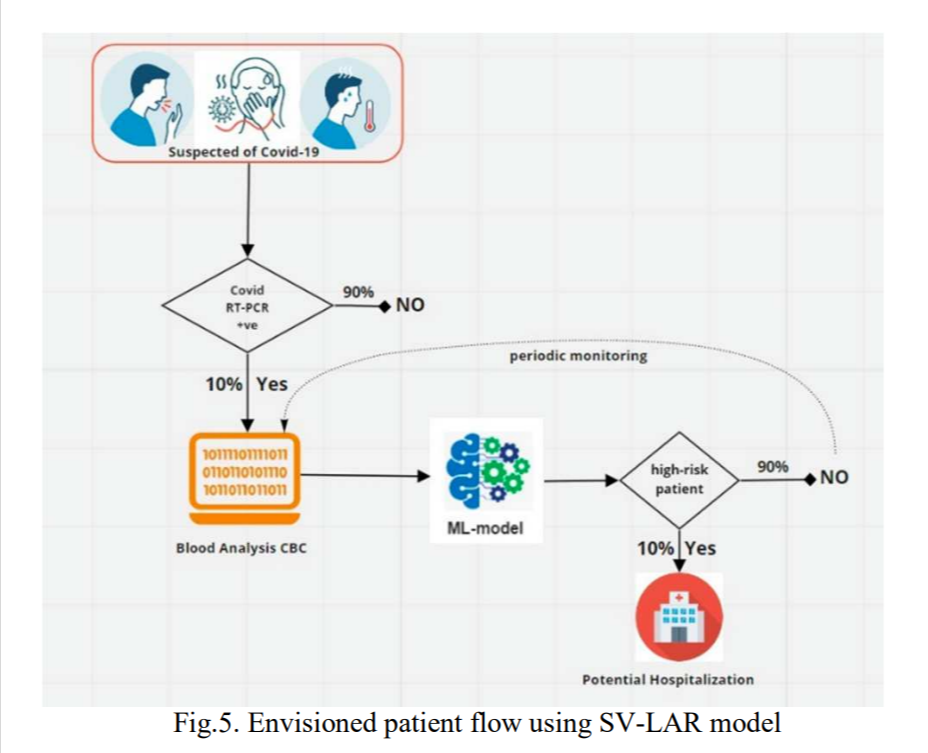
The model predicts their risk of hospitalization. If the patient belongs to high-risk class, the he/she should be tested for other tests like blood gas analysis and should be admitted upon physician approval. If the patient is deemed not at high-risk he/she should be sent back with periodic blood workup. The blood samples should be periodically analysed and tested through the SV-LAR model for risk assessment. This will enable healthcare staff to monitor each patients' development effectively
By the simple intervention of machine learning model (SV-LAR) with f1-score of 87%, the identification of a potential high-risk patient can be performed easily. The differential costs of tests required to the prediction is also underwhelming. SV-LAR model can be used to benefit healthcare workers by identifying about 10% high-risk COVID-19 patients from the increasing COVID-19 patient population. The precision of the model is a high 91% and 83% recall for the positive class. Simply put, 83 out of 100 high-risk patients can be identified correctly using this model and can be taken into hospital care for further treatment.
SV-LAR model is potentially fastest way to triage COVID-19 patients into high-risk and low-risk groups. Not only that, it enables to monitor patients via simple, non-expensive, quick, robust and minimally-invasive blood analysis. The identified high-risk patient population can then be put through more tests and procedures and can be treated accordingly. The low-risk patients, on the other hand, can be remotely monitored for any change in the patients' prognosis via the same model.
Our proposed model can be utilised globally. It relies on basic blood analysis, which is the most simple and established diagnostic service readily available in the healthcare system of any nation. This enables improved management of pandemic agnostic of the socio-economic standing of the nation. An additional positive impact is related to hospital and patient flow management. It allows patient journey to be monitored from a distance providing better isolation of COVID-19 patients. Given that blood samples could be drawn periodically and analysed away from hospital emergency rooms, it allows ERs to work more efficiently despite the pandemic
If RT-PCR and blood analysis samples were to be taken from residence (as this service is available in India, and possibly in other countries as well), many patients can save a trip to the hospital emergency rooms thus saving time for both the hospital staff and themselves. This model can surely help in managing the pandemic related patient flow effectively and efficiently. Also, this will extend care 'as needed' by each patients' condition, where a high-risk patient is not sent home and a 'not at risk patient is managed quickly without risking unnecessary hospital visit. Thus our model ensures using available resource where needed and thereby improves patient outcomes and hospital's healthcare burden as well.
A limitation of our study however, is that not every patient hospitalized needs ICU. Due to the lack of data we could not segment the ICU needing patients from hospitalization requirements. As more data is collected and made available we can further refine the model. We believe that as more data will be incorporated in the model its performance and reliability will increase. At this point, the model is developed from data available from patient emergency room visits from one hospital only. The model remains to be tested across geographies and hospitals for increased robustness. So far we have utilised only machine learning algorithms and not used deep learning algorithms to train the data. One of the reason was the size of the information available. As the available dataset was small we restricted our approach to include machine learning models. It is known that neural networks and deep learning algorithms work better on large datasets or else they tend to overfit. The performance of the model in larger datasets remains to be seen both using proposed machine learning algorithm and transferring similar approach in deep learning algorithms.
REFERENCES
- "Archived: WHO Timeline COVID-19," Who.int. [Online]. Available: https://www.who.int/news/item/27-04-2020-who-timeline---covid-19. [Accessed: 14-Feb-2021].
- "COVID-19 map johns Hopkins Coronavirus resource Center," Jhu.edu. [Online]. Available: https://coronavirus.jhu.edu/map.html. [Accessed: 14-Feb-2021].
- A. Shander et al., "Essential role of Patient Blood Management in a pandemic: A Call for Action: A call for action," Anesth. Analg., vol. 131, no. 1, pp. 74-85, 2020.
- CDC, "COVID-19 and Your Health," Cdc.gov, 03-Feb-2021. [Online]. Available: https://www.cdc.gov/coronavirus/2019-ncov/need-extra-precautions/people-with-medical-conditions.html. [Accessed: 14-Feb-2021].
- Silva, and D. L. Guidoni, "Predicting the disease outcome in COVID- 19 positive patients through Machine Learning: a retrospective cohort study with Brazilian data," bioRxiv, p. 2020.06.26.20140764, 2020.
- W.-J. Guan et al., "Comorbidity and its impact on 1590 patients with COVID-19 in China: a nationwide analysis," Eur. Respir. J., vol. 55, no. 5, p. 2000547, 2020.
- B. Wang, R. Li, Z. Lu, and Y. Huang, "Does comorbidity increase the risk of patients with COVID-19: evidence from meta-analysis," Aging (Albany NY), vol. 12, no. 7, pp. 6049-6057, 2020.
- W. Liu et al., "Analysis of factors associated with disease outcomes in hospitalized patients with 2019 novel coronavirus disease," Chin. Med. J. (Engl.), vol. 133, no. 9, pp. 1032-1038, 2020.
- Y. Zhou et al., "Comorbidities and the risk of severe or fatal outcomes associated with coronavirus disease 2019: A systematic review and meta-analysis," Int. J. Infect. Dis., vol. 99, pp. 47-56, 2020.
- Swiss Society Of Intensive Care Medicine, "Recommendations for the admission of patients with COVID-19 to intensive care and intermediate care units (ICUs and IMCUs)," Swiss Med. Wkly, vol. 150, no. 1314, p. w20227, 2020.
- Z. Zhao et al., "Prediction model and risk scores of ICU admission and mortality in COVID-19," PLoS One, vol. 15, no. 7, p. e0236618, 2020.
- D. Ji et al., "Prediction for progression risk in patients with COVID- 19 pneumonia: The CALL score," Clin. Infect. Dis., vol. 71, no. 6, pp. 1393-1399, 2020.
- E. Grifoni et al., "The CALL score for predicting outcomes in patients with COVID-19," Clin. Infect. Dis., vol. 72, no. 1, pp. 182-183, 2021.
- J.-J. Qin et al., "Redefining cardiac biomarkers in predicting mortality of inpatients with COVID-19," Hypertension, vol. 76, no. 4, pp. 1104 1112, 2020.
- S. Schalekamp et al., "Model-based prediction of critical illness in hospitalized patients with COVID-19," Radiology, vol. 298, no. 1, pp. E46-E54, 2021.
- Y. Zhou et al., "Exploiting an early warning Nomogram for predicting the risk of ICU admission in patients with COVID-19: a multi-center study in China," Scand. J. Trauma Resusc. Emerg. Med., vol. 28, no. 1, p. 106, 2020.
- G. Wang et al., "C-reactive protein level may predict the risk of COVID-19 aggravation," Open Forum Infect. Dis., vol. 7, no. 5, p. ofaa153, 2020.
- S. Chikode, N. Hindlekar, P. Padhye, N. Darapaneni, and A. R. Paduri, "COVID-19: Prediction of Confirmed cases, active cases and health infrastructure requirements for India," International Journal of Future Generation Communication and Networking, vol. 13, no. 4, pp. 2479-2488, 2020.
- Allen Institute For AI, "COVID-19 Open Research Dataset Challenge (CORD-19)."
- P. K. Nigam, "Correct blood sampling for blood gas analysis," J. Clin. Diagn. Res., vol. 10, no. 10, pp. BL01-BL02, 2016.
- "1. Supervised learning scikit-learn 0.24.1 documentation," Scikit-learn.org. [Online]. Available: https://scikit-learn.org/stable/supervised_learning.html. [Accessed: 14-Feb-2021].
- "Welcome to imbalanced-learn documentation! imbalanced-learn 0.7.0 documentation," Imbalanced-learn.org. [Online]. Available: https://imbalanced-learn.org/stable/index.html. [Accessed: 14-Feb-2021].
- K. P. Shung, "Accuracy, precision, recall or F1? towards data science," Towards Data Science, 15-Mar-2018. [Online]. Available: https://towardsdatascience.com/accuracy-precision-recall-or-fl-331fb37c5cb9. [Accessed: 14-Feb-2021].
- "API Reference scikit-learn 0.24.1 documentation," Scikit-learn.org. [Online]. Available: https://scikit-learn.org/stable/modules/classes.html. [Accessed: 14-Feb-2021].
- N. Darapaneni et al., "A machine learning approach to predicting covid-19 cases amongst suspected cases and their category of admission," in 2020 IEEE 15th International Conference on Industrial and Information Systems (ICIIS), 2020, pp. 375-380.
- S. Cantenys-Molina, E. Fernández-Cruz, P. Francos, J. C. Lopez Bernaldo de Quirós, P. Muñoz, and J. Gil-Herrera, "Lymphocyte subsets early predict mortality in a large series of hospitalized COVID- 19 patients in Spain," Clin. Exp. Immunol., no. cei. 13547, 2020.


